To simplify dendritic spine analysis, our histology and IT team joined forces and developed a macro-based image analysis for rater-independent, unbiased, fast, and efficient automated dendritic spine quantification.
Brain sections are stained by the Golgi-Cox method. For spine density analysis, Z-stack images are taken using 63x oil lens, and stacks are collapsed using Zeiss ZEN software. Spines are detected and quantified automatically by a macro running in Image Pro 10 software.
Spine density analyses are of interest for studies associated with different human diseases and their animal models:
- Spine density analysis in the SOD1G93A mouse model of amyotrophic lateral sclerosis. Fogarty et al. 2016. https://doi.org/10.1186/s40478-016-0347-y
- Dendritic spine remodeling accompanies Alzheimer’s disease pathology and genetic susceptibility in cognitively normal aging. Boros et al. 2019. https://doi.org/10.1016/j.neurobiolaging.2018.09.003
- Common defects of spine dynamics and circuit function analyzed in neurodevelopmental disorders, including BTBR T+ tf/J mice as an inbred strain model of autism spectrum disorder. Nakai et al. 2018. https://doi.org/10.3389/fnins.2018.00412
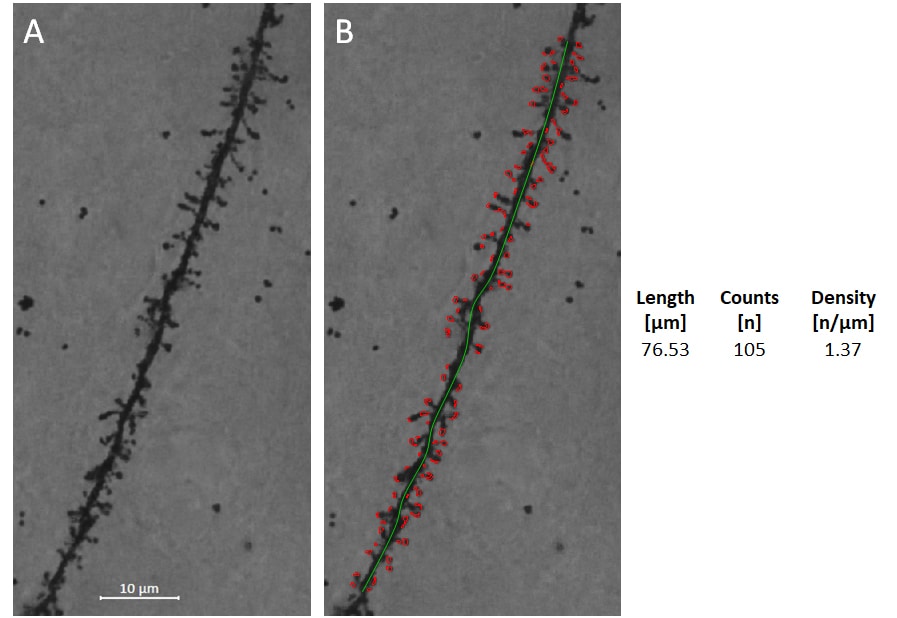
Figure: Spine recognition and spine density analysis. A: Collapsed Z-stack image of Golgi-Cox stained dendrite of frontal cortical pyramidal cell. B: Length of the dendritic segment was determined (green line), and spines were then identified and counted automatically. The data generated from this image are shown on the right.
